Discover, Analyze and Remediate Sensitive Data Anywhere
Personal data where it shouldn't be?
Our fast and accurate on-prem AI will find and fix it.

Control Data Risk with Powerful AI

Discover
Scan for all personal or sensitive information securely. No data leaves your network.

Report
Analyze and review discovered data based on who is affected, where and how.

Remediate
Redact or erase unwanted personal information. Individually or in bulk.

Comply
Keep up with regulations with incremental and scheduled scans.
When to Use PII Tools?
Audits & Compliance
Breach Investigations
Data Migrations
Consultants and MSPs
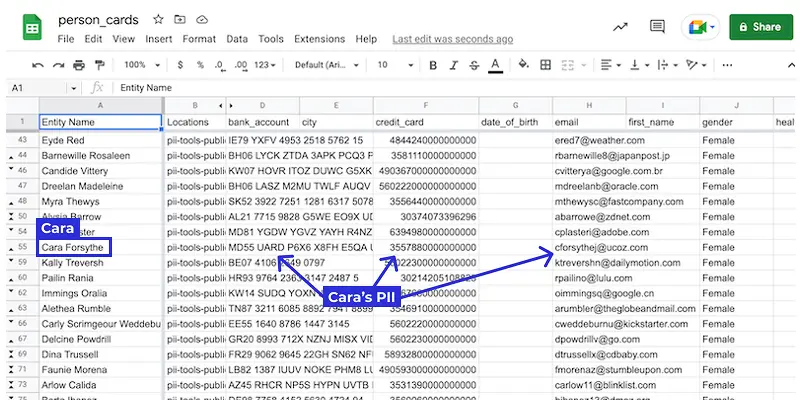
Person Cards® by PII Tools
Document Redaction

AI Data Protector
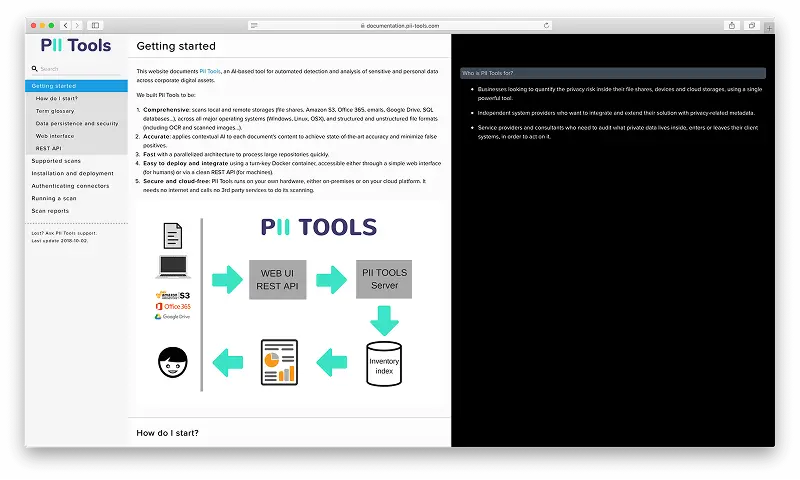
Detect PII Data in 400+ File Formats
PII Tools scans through all the sensitive data, local or cloud-based, structured and unstructured.
What Our Clients Say About PII Tools

Mark Cassetta
SVP Strategy
“Our survey found that 22% of the time, humans failed to identify personal data in documents, while PII Tools succeeded in all scenarios. By integrating with PII Tools, Titus was able to significantly reduce the compliance risk for our customers.”


Raul Diaz
Senior Director, IT
“A manual data review would take us years and years, which was not an option. PII Tools provides us with a full report wherever there is any PII on our Sharepoint, GSuite, Microsoft Exchange, Salesforce, and physical devices.”


Shane Reid
Group Director, CEO – North America
“Integrating PII Tools has allowed Umlaut Solutions to handle petabytes of client data seamlessly, enhancing accuracy and reducing the risk of data breaches.”

Want to know more about PII Tools?
Explore our free resources to see how PII Tools can protect your data and prevent breaches.
Trusted by Top Brands Worldwide
From Purchase to Your First PII Scan in Minutes

Instant Deployment
Install PII Tools on your own server in 30 minutes, using our PII Tools VMware or Docker image.

Scan & Analyze
Discover personal and sensitive data across your digital assets. In motion or at rest.

Take Action
Review, export, or remediate sensitive data from the Analytics dashboard.
Take Control of Your Personal and Sensitive Data Today!
Top 3 Reasons Clients Schedule a Demo with PII Tools:
Avoid or solve data breaches
Comply with GDPR, HIPAA, PCI DSS, and other legislation
Prepare for an audit